This Research Insight covers a study from the Chen Lab. Here, we highlight a proposed model for retinal degeneration based on binding activity of the Cone-Rod Homeobox transcription factor, CRX.

In this study published in eLife, researchers in the lab of Shiming Chen, PhD, Dr. Bernard and Janet R. Becker Distinguished Professor of Ophthalmology at WashU Medicine, investigated the pathogenic mechanisms of mutations in the CRX homeodomain (HD). These findings by Yiqiao Zheng, PhD and colleagues demonstrate the unique mechanisms by which CRX mutations recapitulate human disease due to varied binding interactions with rod and cone photoreceptor genes.
The role of CRX in photoreceptor development and disease
The HD region of a regulatory gene encodes the DNA-binding region of the protein that is responsible for recognizing and binding to specific DNA sites of other genes. This HD-DNA interaction is required for activating the expression of target genes in a process called transactivation. HD-containing transcription factors—such as CRX—are necessary for appropriate organ and tissue development. CRX is expressed by the retina of our eyes, particularly in light-sensing rod and cone photoreceptor neurons. During retinal development, CRX directs the differentiation of rods and cones by activating key photoreceptor genes, which is essential for visual function development. In adults, CRX is required for maintaining vision and for rod/cone cell survival.
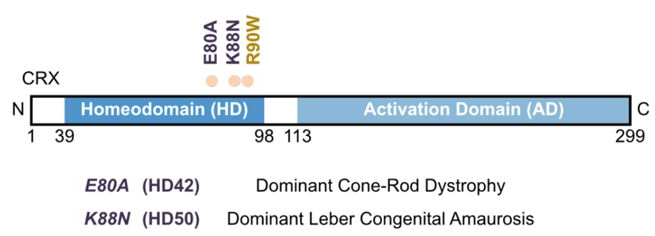
In humans, mutations in the HD region of CRX are linked to severe inherited retinal diseases—notably, autosomal dominant cone-rod dystrophy (adCoRD) and Leber congenital amaurosis (adLCA)—that are characterized by distinct symptoms and progression patterns.
Individuals carrying the CRX HD mutation p.E80A often develop adCoRD, losing color vision and visual acuity, initially due to cone deficits. As adCoRD progresses, rod function also deteriorates, producing blind spots—or scotomas—in their visual field and loss of their peripheral vision. In contrast, individuals carrying a p.K88N mutation often develop adLCA and experience symptoms—including severe vision loss, photophobia, hyperopia, and nystagmus—that begin at birth or during early childhood and often progress to complete blindness within a year.
The ability of mutations to the HD region of CRX to yield such varied clinical presentations is a key area of investigation to understand inherited retinal diseases. To advance this research, scientists in the Chen Lab have developed two mouse models, carrying either the human p.E80A or p.K88N CRX HD mutations. These mutant mice successfully recapitulate key features and progressive vision loss described for human patients diagnosed with adCoRD or adLCA.
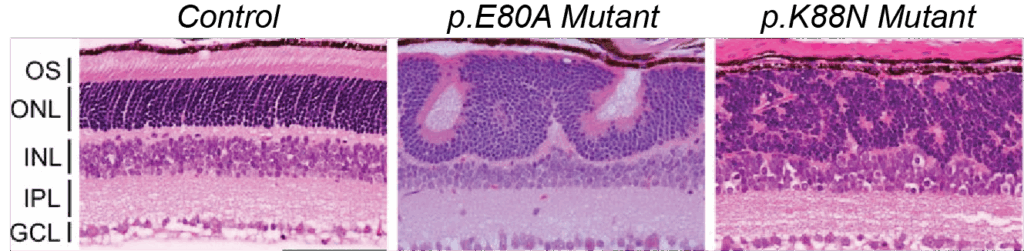
More significantly, in this study, Zheng and colleagues use these mouse models combined with a battery of other tools to uncover two novel mechanisms that drive the retinal pathology associated with dominant CRX HD mutations.
The HD region of CRX has a key role in determining specificity and the effect of CRX-DNA binding
The ability of transcription factors to bind to DNA depends on the local chemistry between their precise amino acid sequence and the highly conserved nucleotide bases in the DNA recognition sites—or consensus motifs—that they bind to. Single amino acid mutations in a transcription factor can therefore alter this local chemistry, potentially changing binding specificity and gene expression.
The specificity of CRX for its consensus motif is determined by the sequence of 60 amino acid residues that comprise the HD region of CRX. Zheng and colleagues hypothesized that the disease-associated mutations in the HD region of CRX—including p.E80A and p.K88N—impact the binding specificity of CRX in unique ways. In turn, the altered CRX-DNA binding could result in distinct changes in developmental gene expression and the drastically different disease phenotypes observed.
Point mutations in the HD region impact CRX-DNA binding ability
Zheng and colleagues developed a series of studies spanning experimental models to assess how p.E80A and p.K88N point mutations in the HD region of CRX impacted DNA binding. They first developed a specific biochemical assay with DNA sequencing as a readout (Spec-seq) to determine the relative binding affinity between (bacterially expressed) recombinant CRX protein and its DNA targets in test tubes. They then compared these results to CRX-DNA interactions in the retinas of normal and mutant mouse models to determine how these p.E80A or p.K88N mutations impact the binding of CRX to target genes and the regulation of their expression.
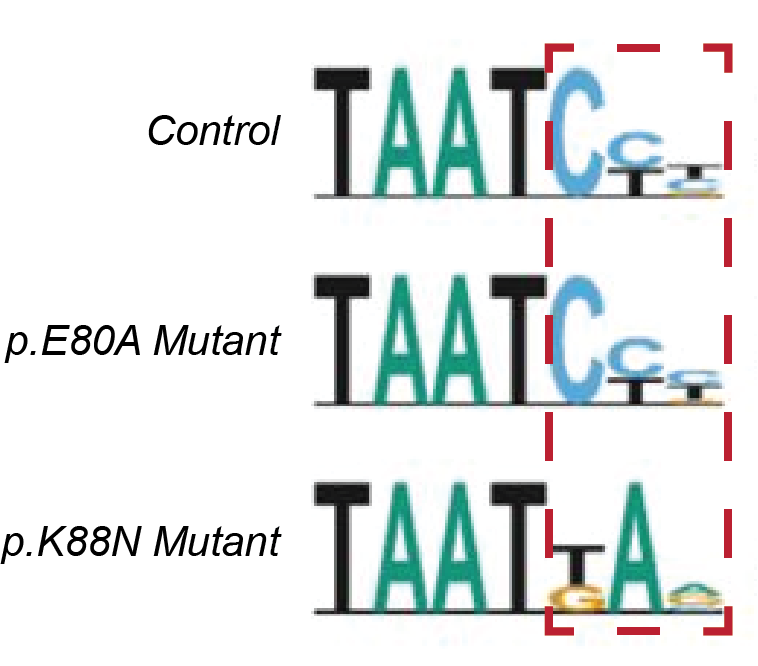
These studies revealed that p.E80A protein bound to the same consensus DNA motifs in photoreceptor genes as the normal CRX in both test tubes and the retina; however, pE80A bound to the DNA with less specificity and/or higher affinity. In contrast, p.K88N protein bound poorly to the canonical CRX consensus motifs but targeted a set of new DNA sites in non-photoreceptor genes. This altered DNA binding specificity indicated the importance of the K88 residue as a specificity determinant. Together, these results discovered that p.E80A and p.K88N impact CRX-DNA binding via distinct molecular mechanisms.
Different point mutations in CRX HD differentially alter gene activation during development
Zheng and colleagues hypothesized that proper CRX-DNA binding triggers precisely timed gene activation to promote photoreceptor differentiation and that CRX HD mutations altering DNA binding will disrupt precise gene activation and photoreceptor functional development. To determine the effects of p.E80A and p.K88N CRX mutations on photoreceptor gene regulation and differentiation, Zheng and colleagues performed a bulk RNA sequencing analysis that compared the transcriptomes of wildtype, p.E80A mutant, and p.K88N mutant developing mouse retinas. This analysis demonstrated that p.E80A and p.K88N mutant proteins altered the timing and/or degree of photoreceptor gene activation during a critical window of development.
The p.E80A and p.K88N mutations yielded opposite effects on gene activation. The p.E80A mutant CRX induced premature upregulation of rod genes and downregulation of cone genes, which led to the adCoRD-like phenotype. In contrast, the p.K88N mutant CRX failed to activate both rod and cone genes, as it improperly bound to non-photoreceptor genes, which completely compromised photoreceptor differentiation. This resulted in an LCA-like phenotype.
Understanding mechanisms of CRX-mediated disease can provide insights into putative gene therapy treatment strategies
Taken together, these findings demonstrate that precise interactions between CRX and its targets are required for healthy retinal gene expression and photoreceptor differentiation. Further, they support a model that E80A and K88N mutations disrupt CRX regulatory activity in unique ways, thereby giving rise to distinct, dominant disease phenotypes that recapitulate human disease.
Understanding pathogenic mechanisms is necessary for tailoring gene therapy strategies to target individual disease phenotypes. These insights can inform the timeline for treatment and choice of gene augmentation with additional healthy CRX copies versus removal of interfering mutant proteins.